Chem. Sci. – George E. Cutsail, Matthew O. Ross, Amy C. Rosenzweig, Serena DeBeer.
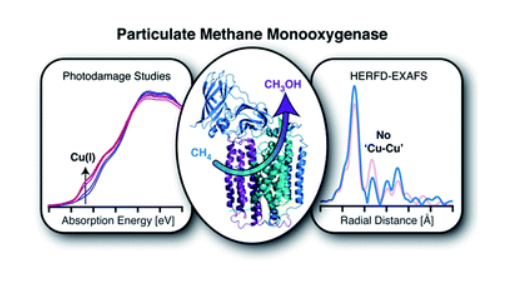
The enzymatic conversion of the greenhouse gas, methane, to a liquid fuel, methanol, is performed by methane monooxygenases (MMOs) under mild conditions. The copper stoichiometry of particulate MMO (pMMO) has been long debated, with a dicopper site previously proposed on the basis of a 2.51 Å Cu–Cu feature in extended X-ray absorption fine structure (EXAFS) data. However, recent crystallographic data and advanced electron paramagnetic resonance (EPR) characterization support the presence of only mononuclear copper sites. To reconcile these data, we have collected high-energy resolution fluorescence detected (HERFD) and partial fluorescence yield (PFY) EXAFS spectra of Methylococcus (M.) capsulatus (Bath) pMMO. Both methods reveal only monocopper sites. These data were compared to previously published pMMO PFY-EXAFS data from M. capsulatus (Bath) and Methylomicrobium alcaliphilum 20Z, supporting dicopper and monocopper sites, respectively. The FT-EXAFS feature previously attributed to a dicopper site can be reproduced by the inclusion of a metallic copper background signal. The exact position of this feature is dependent on the nature of the sample and the percentage of background contamination, indicating that visual inspection is not sufficient for identifying background metallic contributions. Additionally, an undamaged X-ray absorption spectrum was obtained, consistent with the copper oxidation-state speciation determined by EPR quantification. X-ray photodamage studies suggest that the previously observed Cu(I) XAS features are in part attributable to photodamage. This study illustrates the complex array of factors involved in EXAFS measurement and modeling of pMMO and more generally, dilute metalloproteins with multiple metal centers.
Click here for the complete article!